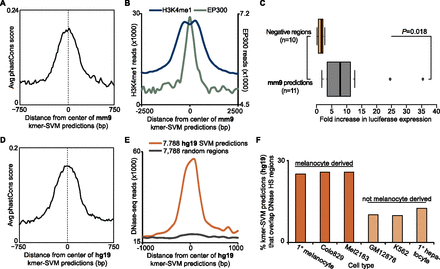
kmer-SVM classifier predicts additional enhancers in the mouse and human genomes. (A–C) Mouse predictions; (D–F) human. (A) Average phastCons score (vertebrate, mm9) in a 1.5-kb window around the centers of the 7361 kmer-SVM-predicted mouse melanocyte enhancers. (B) Number of ChIP-seq reads for H3K4me1 (blue line, left axis) and EP300 (green, right axis) in a 5-kb window around the centers of 7361 predicted melanocyte enhancers (averaged in 100-bp bins). (C) Box plot summarizing results of reporter assays for 10 negative regions (top, orange) and 11 predicted enhancers (bottom, gray). P = 0.01803 by two-tailed t-test. (D) Average phastCons score (vertebrate, hg19) in a 1.5-kb window around the centers of the 7788 kmer-SVM–predicted human melanocyte enhancers. (E) Number of DNase-seq reads from human primary melanocytes in a 2-kb window (averaged in 100-bp bins) around the centers of predicted enhancers (orange) and randomly selected regions matched in size and GC content (gray). (F) Percentage of predicted human melanocyte enhancers (n = 7788) that overlap DNase HS peaks in six cell types, which are either derived from melanocytes (orange) or not (beige).











