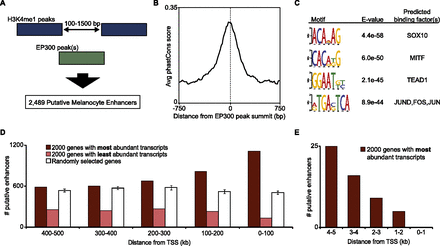
H3K4me1-flanked EP300 peaks have multiple characteristics of melanocyte enhancers. (A) Visual representation of our approach to identify putative melanocyte enhancers (described in text and Methods). (B) Average phastCons score (vertebrate, mm9) in a 1.5-kb window around the summit of 2489 putative melanocyte enhancers. (C) Top four motifs enriched in sequences of putative enhancers, with corresponding E-values (enrichment P-value times number of motifs tested; calculated by DREME) and factors predicted to bind to these motifs. (D) Number of putative enhancers in 1-MB window (100-kb bins) around the TSS of 2000 genes with the most abundant transcripts in melan-a (dark red), 2000 genes with least abundant transcripts in melan-a (light red), and 2000 randomly selected genes (white; average of five sets with SD represented by error bars). (E) Similar analysis to D, but using a 10-kb window and 1-kb bins.











