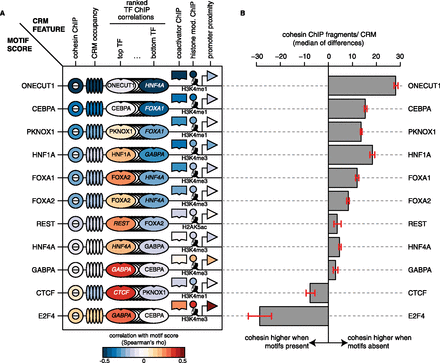
Cohesin ChIP signal is significantly associated with TF motif score. (A) Cartoon heatmap representation of correlations between each sequence-specific factor's motif score and the ChIP signal of all available ChIP-seq data sets. Correlations with CRM occupancy (number of distinct TFs present) and promoter proximity (distance to the nearest canonical TSS) are also shown. For each factor, the motif score correlation was calculated on the set of CRMs that contained a ChIP-seq peak for the same factor. Correlations with cohesin and coactivator ChIP signal were averaged over subunits (RAD21, STAG1, STAG2) and family members (EP300, CREBBP), respectively. Heatmap rows were ordered by increasing correlation with cohesin ChIP signal (from top to bottom). As a visual summary, only the top- and bottom-ranking correlations involving TFs are shown (see Supplemental Figs. S6, S7 for all correlations). (B) Increased cohesin ChIP signal at TF binding events without motifs. For each sequence-specific factor, the number of cohesin ChIP fragments within CRMs without high-scoring motifs was compared with that of CRMs with motifs. The 95% confidence intervals shown are based on a normal approximation of the Hodges–Lehmann estimate (median of all possible differences).











