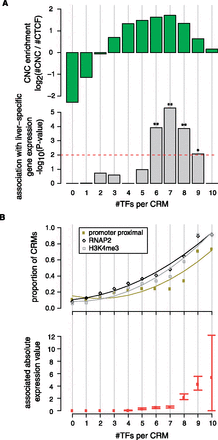
CNC sites are associated with liver-specific gene expression. (A) Ratio of CNC-containing CRMs versus those with CTCF (log-fold change) for CRM classes with 0–10 TFs. Each class of CRMs was also tested for association with 107 genes signficantly up-regulated in mouse liver cells (see Methods). The significance of the association (negative-log-transformed Fisher's exact test P-values) are indicated. (*) P < 0.01; (**) P < 0.001. The enrichment of CNC-containing CRMs reaches threefold when seven TFs are present, and coincides with highly significant enrichment for an association with liver-specific gene expression for the same class. (B) CRMs with high numbers of colocalizing TFs are associated with increased promoter proximity (≤2.5 kb from an annotated TSS) and characteristics of transcriptional activity (RNAP2 and H3K4me3 ChIP-seq peaks). Likewise, the associated absolute gene expression value increases significantly with the number of bound TFs. Error bars indicate the 95% confidence interval of the median.











