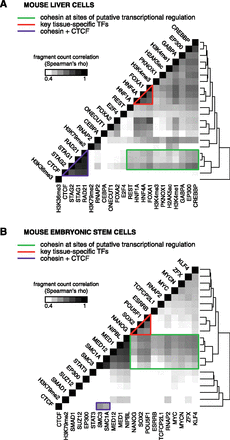
Within-CRM binding correlations reveal distinct modes of cohesin binding in diverse cell types. The number of ChIP fragments (mapped reads extended to the estimated fragment length) overlapping a given CRM was used as a measure of binding strength for each data set. Factors were clustered along both axes based on the similarity in their colocalization profiles. (A) Heatmap visualization of all pairwise correlations between all ChIP-seq data sets in mouse liver cells illustrates cohesin subunits (RAD21, STAG1, STAG2) clustered with CTCF. Cohesin also correlates with key tissue-specific TFs (FOXA1, HNF4A, and HNF1A) independently of CTCF as well as with histone modifications associated with transcriptional activity (H3K4me1, H3K4me3, H2AK5ac) and coactivators (EP300 and CREBBP). (B) All pairwise correlations between previously published ChIP-seq data sets in mouse embryonic stem cells. Cohesin binding strength (SMC1A, SMC3) correlates with CTCF while also forming a distinct cluster with key regulators of stem cell identity (POU5F1, SOX2, NANOG, MYC), components of the mediator complex, as well as RNAP2. Similar results were obtained by performing the correlation analysis separately on CRMs with CNC and CTCF (see Supplemental Fig. S3).











