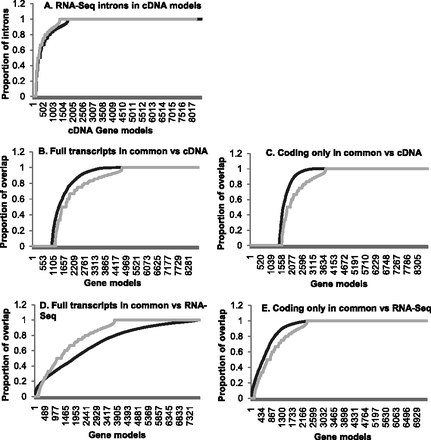
Intron, exon, and base coverage of cDNA-supported Ensembl gene models. The full transcript, and if available, corresponding protein-coding regions of 8822 cDNA-supported Ensembl models were extracted from Ensembl version 60. (A) All of the introns from the 8504 multiexon full transcripts were compared with the RNA-seq intron database and scored positive if there was an exact match. The proportion of exact match introns was calculated for each transcript individually and plotted (dark gray). The process was repeated for the protein-coding portion of 8227 multiexon gene models (light gray). (B) The 8822 cDNA-supported Ensembl models were compared with the RNA-seq models, and the single best overlap model was identified. The proportion of intersecting nucleotides (dark gray) and exons (light gray) compared with the cDNA-supported models was calculated for each gene model and plotted. (C) The same calculation as in B using only the protein-coding regions. (D) The 7801 cDNA-supported Ensembl models that overlap an RNA-seq model were compared. The proportion of intersecting nucleotides (dark gray) and exons (light gray) compared with the best overlap RNA-seq model was calculated for each gene model and plotted. (E) The same calculation as in D using 7386 cDNA-supported models where the best match full transcript was from the same RNA-seq model as the best match coding region. Exon matching in B–E did not include the transcription start or stop.











