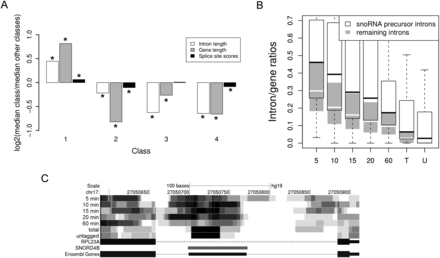
Properties of intron classes and snoRNA processing. (A) Log2 fold-changes of the median in each class compared with the median of all other classes are shown for intron length (white), gene length (gray), and splice site scores (black), calculated using MaxEntScan (Yeo and Burge 2004). (*) Statistically significant difference compared with all other classes (Wilcoxon test, FDR corrected P-value ≤ 0.05). With the exception of splice site scores in Class 3, distributions showed significant differences for all classes and all features analyzed. (B) Intron/gene ratios are shown separately for snoRNA precursor introns (white) and all other introns (gray). Intron/gene ratios for precursor introns were significantly increased compared with all other introns at all times. (C) Read density plot illustrating the processing of an exemplary precursor intron to the mature snoRNA. One can observe both the substantially slower decay of the intronic sequences surrounding the mature snoRNAs, which was typical for intronic snoRNAs, and the characteristic 5′ read start pattern of mature snoRNAs in total and untagged RNA.











