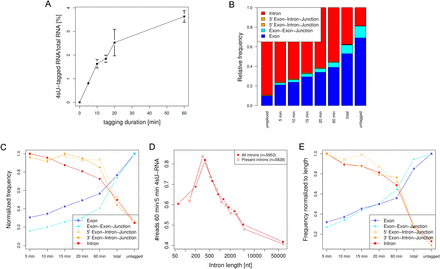
Ultrashort and progressive 4sU-seq reveals the kinetics of RNA splicing. (A) Contribution of newly transcribed RNA to total RNA levels following different durations of 4sU exposure. The relative contributions of purified newly transcribed RNA (4sU-tagged RNA) to input RNA (total RNA) measured spectrophotometrically (OD260) are shown (combined data of three biological replicates). Standard error is indicated by error bars. (B) Distribution of the number of reads mapped to exons, exon–exon junctions, exon–intron junctions, and intron regions for 5- to 60-min 4sU-RNA, total and untagged RNA. Samples are ordered according to increasing age of RNA in these samples from 5- to 60-min 4sU-tagging to total RNA and finally untagged RNA. RNA in the untagged RNA samples is at least 60 min old. This visualizes the maturation of transcripts over time. The expected distribution of reads for completely unspliced RNA is shown in the left-most column (see Supplemental Methods). (C) Normalized read frequencies were calculated by first dividing read numbers by the total number of reads on protein-coding genes in the corresponding sample. Subsequently, frequencies for a specific read type were divided by the maximum frequency observed for the corresponding read type in any sample. Despite the overall smaller number of exon–exon junction and exon–intron junction reads, their normalized read frequencies are similar to normalized frequencies of exon and intron reads, respectively. (D) Introns were binned into 10 equally sized groups according to intron length. For each group, average ratios of intron read counts in 60-min compared with 5-min 4sU-RNA were calculated. Results are shown both for all introns of the 525 most highly expressed genes (excluding only those with 0 reads in 5-min 4sU-RNA) and introns present in 5-min 4sU-RNA (RPKM ≥ 0.5). Ratios were highest for introns in a range of 300–400 nt and decreased both for longer and shorter introns. This was observed independently of gene length (see Supplemental Fig. 1E). (E) After normalizing the number of intronic reads to intron length and exonic reads to exon length, the delay between the decrease in intronic reads and exon–intron junction reads, on the one hand, and the increase in exonic reads and exon–exon junction reads, on the other hand, disappeared. For this analysis, only the 525 most highly expressed genes were considered. After normalization to intron and exon length, respectively, the same normalization was applied as in C.











