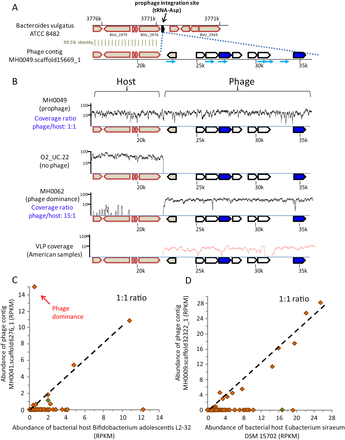
Lysogenic lifestyles of gut microbiota phages. (A) The 5′ end of phage contig MH0049.scaffold15669_1, which was assembled in sample MH0049 (Danish individual), has a 99.5% identity in the sequenced genome of Bacteroides vulgatus ATCC 8482. (Block arrows) Genes; (cyan-colored arrows) spacers matching the phage contig. (B) Coverage of phage contig MH0049.scaffold15669_1 by MetaHIT metagenomic reads from three samples. (X-axis) Position on phage contig; (y-axis) read coverage (log scale). (Red curve) Coverage of VLP reads from Reyes et al. (2010). (C,D) Abundance of phages MH0041.scaffold6276_1 and MH0009.scaffold32322_1 and their respective bacterial hosts in MetaHIT samples, indicative of lysogeny as a preferred lifestyle. The x- and y-axes represent abundance of host and phage, respectively. Each data point represents a European individual sampled as part of the MetaHIT gut microbiota project (Qin et al. 2010). Green-colored samples are the ones in which a spacer matched the phage sequence. “Phage dominance” indicates samples where the phage is suspected to have become active. The correlation coefficient of phage and host abundances for samples where both existed is 0.4 and 0.98, respectively.











