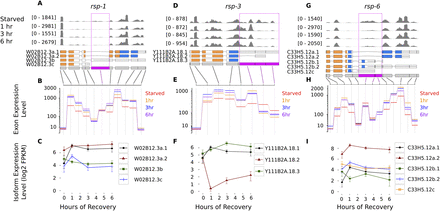
Genes encoding SR (serine/arginine rich) proteins are regulated by alternative exon expression during recovery from starvation. Each column shows the mean coverage of reads over the gene model for each condition (A,D,G), the fitted expression for each exon from the generalized linear model used by DEXSeq (B,E,H), and Cufflinks' estimate of the expression level of each isoform in the gene model (C,F,I). To highlight relative changes in exon expression, gene browser coverage is scaled to the maximum coverage for the gene. Exons identified by DEXSeq as being alternatively expressed (FDR 5%) are highlighted in magenta in the gene model. Transcript sequence annotated as “RNA-recognition motif” by Pfam is highlighted in orange in the transcript models. Transcript sequence rich in serine and arginine (the SR proteins' RS domains) is highlighted in blue. (Dotted line) Untranslated regions. Error bars on Cufflinks' estimates of isoform levels are 95% confidence intervals of the mean. All genes are plotted with their 5′ ends to the left, regardless of strand.











