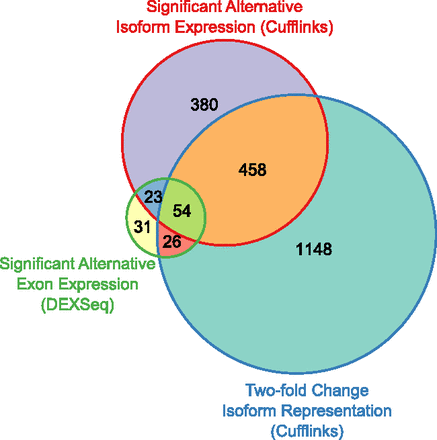
Overlap between three metrics for identifying alternative isoform expression. Genes identified by DEXSeq with alternative exon expression are generally identified by Cuffdiff with significant alternative isoform expression. However, nearly half of the genes identified by Cuffdiff have less than a twofold change in isoform expression. The set of genes identified by Cuffdiff with alternative isoform expression is the union of genes with a significant change in CDS, protein-coding domains, or 3′ UTR. Likewise, the set of genes with twofold change in fractional isoform representation is the union of genes with twofold change in their representation in any of those three categories. The set of genes identified by DEXSeq with alternative exon expression (FDR 5%) is the result of a single test over the entire time series. The set of genes used to make the diagram is the intersection of genes tested by both Cuffdiff and DEXSeq for alternative isoform or exon expression, respectively.











