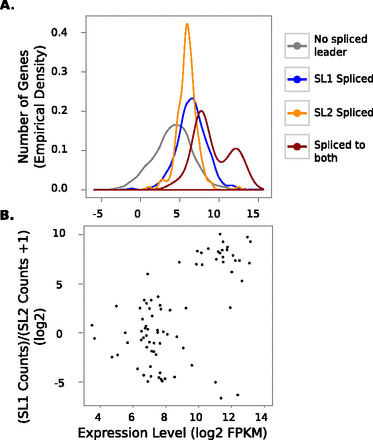
The expression level of genes with transcripts spliced to both SL1 and SL2 reads is bimodal and correlates with the ratio of SL1/SL2 reads. Data for the first hour of recovery are plotted, but this pattern is also present in all other time points (data not shown). Correlation between high expression and high SL1 trans-splicing in addition to SL2 splicing suggests that the activity of an internal promoter in addition to the operon promoter contributes to expression of these most highly expressed genes. (A) The empirical density function of the expression level of transcripts trans-spliced to SL1, SL2, neither, or both is plotted. Trans-spliced transcripts are more highly expressed than transcripts without trans-spliced leaders. (B) The correlation of gene expression with the ratio of SL1/SL2 is plotted for the set of genes with transcripts trans-spliced to both SL1 and SL2. A transcript is considered to be trans-spliced to a particular trans-spliced leader if it received more than 10 reads bearing its sequence. Error bars are 95% confidence intervals of the mean. Of the genes, 2488 genes are in the “no spliced leader” group, 1045 are in the SL1 group, 184 are in the SL2 group, and 80 are in the “both” group.











