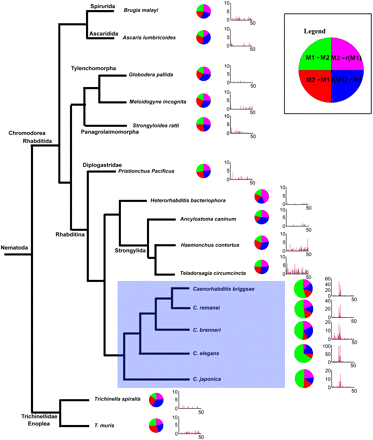
Evolutionary conservation of the motif pair in genomic sequences of 17 nematode species. All occurrences of the two motifs with distance <100 between them were considered. The distributions of the four possible arrangements are shown in pie charts. Large pie charts indicate a statistically significant large number of occurrences of M1 → M2. A histogram of the distances between the two motifs in occurrences of M1 → M2 is shown next to each pie chart (only distances <50 are shown). All Caenorhabditis genomes contain a large number of occurrences of M1 → M2, nearly always with 16–20 bases between the two motifs. All of the other nematodes we analyzed do not exhibit these phenomena. For tree construction we used the published phylogenies in De Ley (2006), Kiontke et al. (2007), and Jex et al. (2010).











