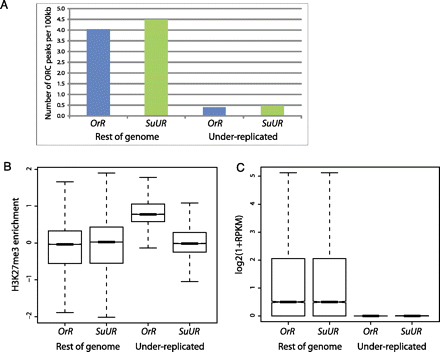
Quantitative comparisons of ORC binding, H3K27me3 enrichment, and RNA expression levels between Oregon-R (OrR) and SuUR mutant in under-replicated regions and the remainder of the genome. Note that in the SuUR mutant these regions are no longer under-replicated, but we refer to them as UR. (A) Comparison of number of ORC binding sites per 100 kb of genome, as assayed by ChIP-seq, shows that in both OrR and SuUR mutants the under-replicated regions have many fewer ORC binding sites than the remainder of the genome. (B) Boxplots showing quantile-normalized H3K27me3 ChIP-chip data, comparing level of H3K27me3 in under-replicated regions with the rest of the genome, in OrR and SuUR mutant salivary glands. OrR UR was significantly enriched for H3K27me3 over the SuUR mutant UR with a P-value < 1 × 10−15 (unpaired Wilcoxon test). (C) Boxplots showing quantile-normalized RNA sequencing data (figure from one replicate; second is indistinguishable) comparing transcriptional levels in under-replicated regions with the rest of the genome in OrR and SuUR mutant salivary glands. Mutation of SuUR does not restore transcription in UR regions. In both strains, the transcription levels in the under-replicated regions were significantly lower than the remainder of the genome (P < 1 × 10−15, Wilcoxon test). The difference between the under-replicated regions in the two strains was not statistically significant (P = 0.3234, Wilcoxon test).











