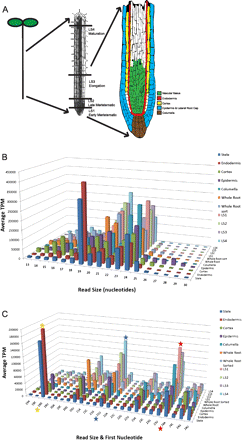
Small RNA characterization in the Arabidopsis thaliana root. (A) Overview of the Arabidopsis root developmental zones and cell types used in this study. (From left to right) Schematic drawing of an Arabidopsis seedling; the root denoting developmental zones isolated by hand sectioning; and the root tip displaying the cell types analyzed in this study. The colors indicate the regions covered by the GFP marker lines used for isolation of cell types by cell sorting. (B) Size distributions display preferences for small RNAs of different types. Size distribution of the reads in the radial and longitudinal data sets is shown. Reads were normalized to transcripts per million (TPM), and the two biological replicates of each radial cell type were averaged. (C) Highly abundant small RNA species are reflected in the read size and the identity of the first nucleotide. Read size by first nucleotide of the reads from the radial and longitudinal data sets is depicted. Canonical miRNAs are 21 nt and begin with a U (blue star), while heterochromatin-associated siRNAs are 24 nt and begin with an A (red star) (Mi et al. 2008). The 19-nt peak corresponds to tRNA fragments, similar to what was reported in roots (orange star) (Hsieh et al. 2009).











