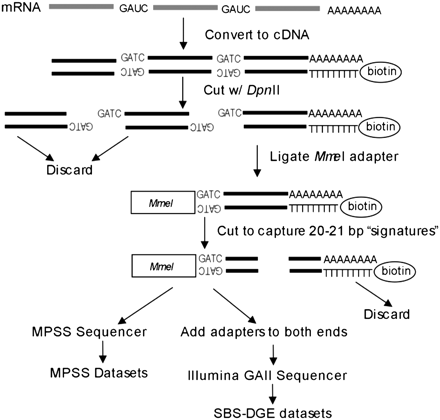
Sample preparations steps for MPSS-DGE and SBS-DGE sequencing. mRNAs were isolated and reverse-transcribed into cDNA, with a biotin tag attached to an oligo-d(T) primer. This was digested with the DpnII enzyme, and only the cDNA fragment closest to poly(A) tail is retained for the next step. An MmeI adapter was added to the 5′ end of cDNA fragments, and the resulting fragments were digested with MmeI, which cuts 20–21 bp downstream from the recognition site, to generate 20–21-bp short fragments called signatures. All signatures were then sequenced by MPSS (Meyers et al. 2004b) or SBS using an Illumina GAII sequencer (for SBS-DGE) (Venu et al. 2011) after adding sequencing adapters on both ends.











