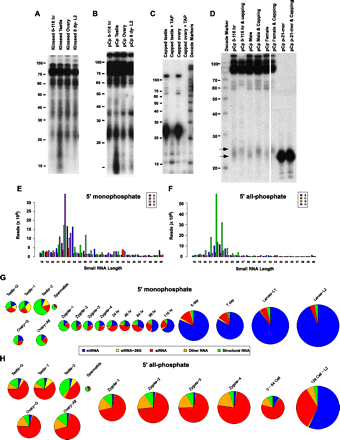
Characteristics of Ascaris small RNAs. (A) 5′ end-labeled Ascaris small RNAs. Low-molecular-weight (LMW) enriched RNAs were treated with calf alkaline phosphatase and then 5′ end labeled with 32P using T4 polynucleotide kinase. RNAs in A–D were resolved on 12.5% denaturing PAGE and autoradiographed. (B) 3′ end-labeled Ascaris small RNAs. LMW enriched RNAs were 3′ end-labeled using 32pCp and T4 RNA ligase. (C) Ascaris small RNAs ∼22 nt in length can be cap-labeled. LMW enriched RNAs were capped using 32P-α-GTP and vaccinia guanylyltransferase. Samples were treated with tobacco acid pyrophosphatase (TAP) to remove the cap to confirm cap-labeling. Note the significant labeling of RNAs ∼22 nt in length. (D) The majority of germline and early embryo 22-nt RNAs have 5′ polyphosphates. pCp-labeled small RNAs shown in B were treated with GTP and vaccinia guanylyltransferase. The lower and upper arrows mark the average position of the labeled small RNAs prior to and after capping. Note that the majority of the 32pCp-labeled RNAs are shifted to a longer length. (E,F) Size distribution of analyzed Ascaris small RNAs. (E) All 28 5′ monophosphate libraries characterized combined. (F) All 19 5′ all-phosphate (5′ tri-, di-, and monophosphate) libraries characterized combined. In these and all subsequent size distribution figures, small RNAs starting with A, C, G, and U in different sizes are plotted against their read frequencies (raw reads, before normalization). (G,H) Comparison, classification, and levels of Ascaris small RNAs characterized. (G) 5′ monophosphate RNAs. (H) 5′ all-phosphate libraries derived from different developmental stages. Area of the circles is proportional to the amount of small RNAs present in each stage or tissue after normalization (see text).











