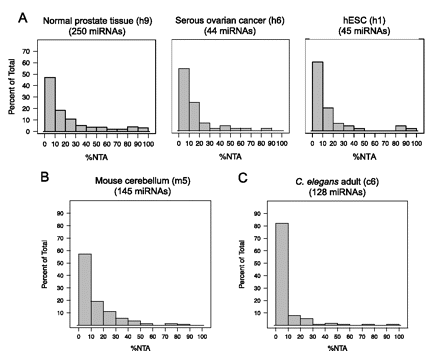
3′ Nontemplated nucleotide additions are miRNA specific. (A) Histograms of %NTA observed in three representative human miRNA-sequencing data sets (normal prostate tissue [h9], serous ovarian cancer [h6], and human embryonic stem cells [h1]). MicroRNAs were required to have 10 or more reads in a given data set to be included, and the number of miRNAs that qualified is given in parentheses for each histogram. The y-axis scale is the same for all three human histograms. (B,C) Histograms of %NTA for representative data sets corresponding to mouse cerebellum or medulloblastoma (B) (m5 in Supplemental Table S1) and the L1 developmental stage of C. elegans (C) (c6 in Supplemental Table S1). MicroRNAs were required to have 10 or more reads in a given data set to be included. Data set identifiers are provided in parentheses for all histograms and refer to descriptions of these data sets provided in Supplemental Table S1.











