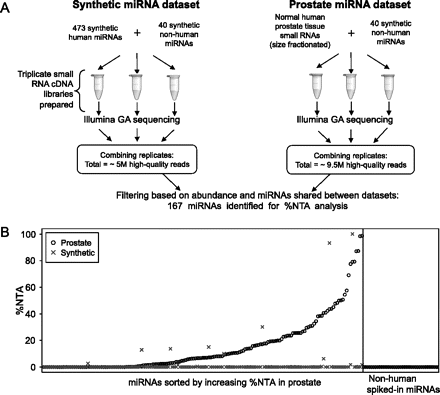
%NTA observed in Illumina Genome Analyzer sequencing of a pool of chemically synthesized miRNAs or of prostate-tissue-derived miRNAs. (A) The schematic describes the workflow for the generation and sequencing of triplicate cDNA libraries corresponding to a pool of chemically synthesized miRNAs and normal human prostate tissue. (B, left) The %NTA for 167 shared miRNAs present at 10 or more reads in both the chemically synthesized miRNA and prostate sequencing data sets are plotted along the y-axis. Normal prostate tissue data are plotted as (○) and the synthetic miRNA data as (×). MicroRNAs are sorted along the x-axis with respect to increasing %NTA observed in prostate-tissue sequencing data. Only seven of the 167 miRNAs analyzed demonstrated greater %NTA in the synthetic miRNA pool compared with its prostate tissue %NTA. In most cases, these represented an additional nucleotide identical to the terminal nucleotide expected for the canonical miRNA sequence (data not shown), which is consistent with a typical type of error expected with solid-phase RNA oligonucleotide synthesis. (Right) The %NTA for 40 nonhuman (a mix of plant and C. elegans) spiked-in miRNAs is plotted for both normal prostate tissue and the synthetic miRNA sequencing data sets.











