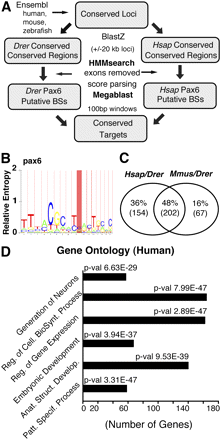
Bioinformatics approaches used to identify the PAX6 target sites and the initial analysis of results. (A) Flow diagram of the HMM approach used to identify evolutionarily conserved direct targets of pax6. The procedure to identify pax6 target enhancers and associated genes is based on the screen of evolutionarily conserved noncoding genomic sequences from orthologous loci in zebrafish and human (or mouse). (B) The screens were performed using models based on pax6 binding sites or their respective reverse complement sequences. (C) 48% (202) of the putative target genes were found in both pairwise comparisons, while 16% (67) are mouse/zebrafish and 36% (154) human/zebrafish specific. (D) The full set of human conserved target genes was analyzed using gene ontology. The identified target genes are enriched for transcription factor activity and developmental regulation as well as for neuron generation functions. The x-axis represents the number of genes for each of the highest over-represented molecular function GO term which are named on the y-axis, and the associated P-values are shown.











