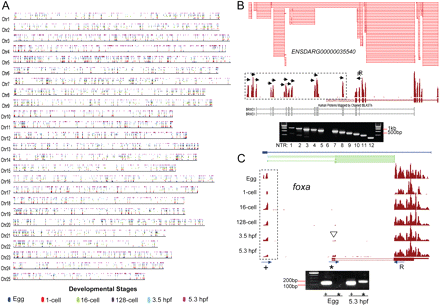
Discovery of NTRs. (A) Genome-wide distribution of NTRs plotted at their mapped chromosomal positions. The color code depicts at which stage an NTR was detected. (B) Novel exons of ENSDARG00000035540, some of which correspond to exons of its human homolog, BRWD1. Splice junctions were also identified among these novel exons. RT-PCR performed using reverse primer at annotated exon 2 (R) and forward primers at each of the 12 NTRs (inset) showed that at least 11 of these NTRs are expressed and are part of the transcription unit. (C) Novel exon in foxa found 5′ of annotated transcription start site. Note the presence of the 5′ annotated exon at 3.5 hpf and 5.3 hpf (white arrowheads). Di-tag (blue horizontal lines) provides support for an isoform that includes the novel exon. Junction mapping at 5.3 hpf (green horizontal lines) indicated the presence of two distinct transcript isoforms. RT-PCR using a reverse primer (R) designed on the 3′ distal exon and forward primers on either 5′ annotated (*) or NTR (+) confirmed the existence of a new 5′ distal exon (inset). The 5′ annotated exon (*) is skipped in the maternal mRNAs (egg to 128-cell stage).











