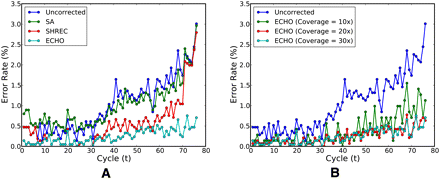
Position-specific by-base error rates for 76-bp PhiX174 data D1. (A) Error rates before and after applying the three error-correction algorithms for sequence coverage depth 30. Spectral alignment (or SA) (see Chaisson et al. 2009) and SHREC (Schröder et al. 2009) are able to improve the error rate in intermediate positions, but they both become less effective for later positions. In contrast, ECHO remains effective throughout the entire read length, reducing the error rate at the end of the read from about 5% to under 1%. (B) Error rates before and after running ECHO with varying coverage depths. ECHO's ability to correct sequencing errors improves as the sequence coverage depth increases. A coverage depth of 15 seems sufficient to control the error rate throughout the entire read length.











