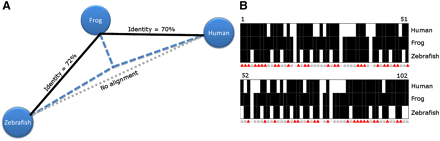
Conservation tunneling. (A) Phylogenetic tree constructed for three orthologous sequences in human (hg18: chr18:53271349–53271555), frog (xenTro2: scaffold_97:133388–133595), and zebrafish (danRer5: chr24:28243171–28243307). Only the human and the frog sequences and the frog and the zebrafish sequences can be aligned (with at least 70% identity across at least 100 bp). The frog sequence has evolved more slowly relative to the human and zebrafish sequences, and thus, can be used to establish the orthology of the diverged human and the zebrafish sequences. (B) Pairwise sequence comparisons. Eighty-seven percent of frog nucleotides are conserved in either human or fish (gray squares), while only 42% are conserved in both human and fish (red triangles).











