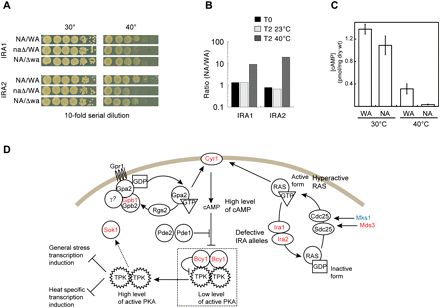
IRA1 and IRA2 are high temperature growth QTLs. (A) Reciprocal hemizygosity confirms that IRA1 and IRA2 are high temperature growth QTLs. WA/NA hybrids were individually deleted for the IRA alleles and used to assess their contribution to high temperature growth. Plate spotting assay using 10-fold serial dilution demonstrates better growth of the hybrid when the NA allele is present. (B) Competition experiment on hybrids with IRA1/IRA2 reciprocal hemizygous deletions (such as A) that resembles the selective step applied to the pool. Hybrids carrying the NA allele outcompete ones with WA allele after 192 h (T2) of growth at 40°C. (C) Internal level of cAMP is reduced at 40°C, but unchanged at 30°C for WA/NA hybrids with WA alleles deleted at both IRA1 and IRA2 loci, compared to NA alleles deleted. (D) RAS/cAMP signaling contributes to natural variation in heat sensitivity. Defective function of the WA alleles of IRA1 and IRA2 at high temperature results in hyperactive RAS, leading to high level of cAMP and high PKA activity inhibiting the heat transcription induction. As a response to heat stress, the majority of the QTLs selected in the pool are from the NA genetic background (red: NA; blue: WA). Dashed arrow indicates unknown mechanism. Figure adapted from Figure 2 of Santangelo (2006) and reprinted with permission from the American Society for Microbiology.











