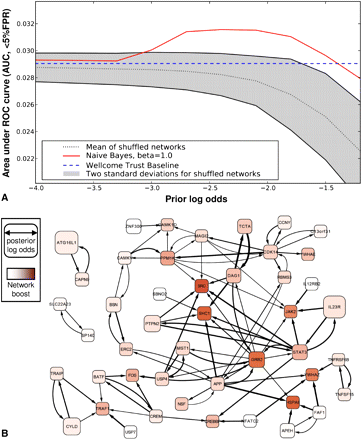
Consideration of the human gene network boosts recovery of validated Crohn's disease genes from GWAS analysis of 2000 cases and 3000 controls. (A) The performance improvement achieved by network-boosted GWAS relative to GWAS alone (Wellcome Trust Baseline, [Wellcome Trust Case Control Consortium 2007]), measuring performance as the area under a ROC curve up to 5% false positive rate (AUC, <5% FPR) for recovering the top 22 Crohn's disease genes identified in a larger meta-analysis of 4549 cases and 5579 controls (Barrett et al. 2008). For the AUC (<5% FPR) measure of performance, a perfect predictor achieves a score of 0.05, while random predictors score near 0.00125. The network boosted approach (colored red line) outperforms the GWAS alone (straight dashed blue line) over a wide range of parameter values. For comparison we also show the results of network boosting when randomized networks are used, plotting the mean (dotted line) and range of performance (2 SD) for 1000 random trials. B plots the network of candidate genes (rounded rectangles) identified from the combination of HumanNet and GWAS data, visualized using Cytoscape (Cline et al. 2007). The node size corresponds to the strength of the combined evidence from the Wellcome Trust Case Control Consortium (WTCCC) data and the network, and the intensity of the red color indicates how much the gene was boosted by the HumanNet GBA. HumanNet linkages are drawn as directed arrows connecting genes, with edge weight scaled by strength of boost contributed by the source to the sink. All genes are drawn with positive posterior log-odds when the prior log-odds of association are −1.7, except for network singletons, and the 50 highest scoring nonsingleton genes are shown. Note the strong boost given to GRB2 and SHC1, which are known to be involved in healing gastric ulcers (Pai et al. 1999), and to JAK2 and STAT3, which were also identified in later meta-analyses (Van Limbergen et al. 2009).











