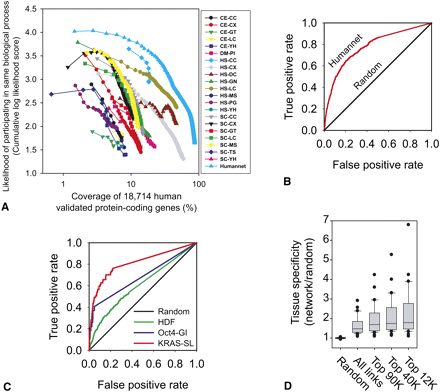
Construction and evaluation of a genome-scale human gene network, HumanNet. (A) 21 diverse functional genomic and proteomic data sets (Supplemental Table S1) were evaluated for their tendencies to link genes in the same biological processes. Pairwise gene linkages derived from the individual data sets were then integrated into a composite network of higher accuracy and genome coverage than any individual data set. The integrated network (HumanNet) contains 476,399 functional linkages among 16,243 (86.7%) of the 18,714 genes encoding validated human proteins. The x-axis indicates the log-scale percentage of the 18,714 genes covered by functional linkages derived from the indicated data sets (curves); the y-axis indicates the predictive quality of the data sets, measured as the cumulative log likelihood of linked genes to share Gene Ontology (GO) biological process annotations, tested using 0.632 bootstrapping and plotted for successive bins of 1000 linkages each (symbols). Data sets are named as XX-YY, where XX indicates species of data origin (CE, C. elegans; DM, D. melanogaster; HS, H. sapiens; SC, S. cerevisiae) and YY indicates data type (CC, co-citation; CX, mRNA coexpression; DC, domain co-occurrence; GN, gene neighbor; GT, genetic interaction; LC, literature-curated protein interactions; MS, affinity purification/mass spectrometry; PG, phylogenetic profiles; PI, fly protein interactions; TS, tertiary structure; and YH, yeast two-hybrid). Detailed descriptions are listed in Supplemental Table S1. (B) Essential genes were highly interconnected in HumanNet, and thus predictable from the network, as shown by ROC analysis. Genes were ranked by their sum of network edge weights to the known essential genes, measuring recovery of known essential genes (true positives) and other genes (false positives) using leave-one-out cross-validation. (C) Genes involved in more specific cellular phenotypes—host factors required for HIV infection (HDF) (Brass et al. 2008), modulators of OCT4 (also known as POU5F1) expression (Oct4-GI) (Ding et al. 2009), and synthetic lethal partners of activated KRAS alleles (KRAS-SL) (Luo et al. 2009)—were also well predicted by their interconnectivity in HumanNet, calculated as for B. (D) Finally, network-linked gene pairs were substantially more likely to show similar tissue specificity in their expression patterns, measured as the likelihood of co-occurrence of transcripts of pairs of genes in the same tissues across 30 different human tissues from the TiGER database of tissue-specific gene expression and regulation (Liu et al. 2008).











