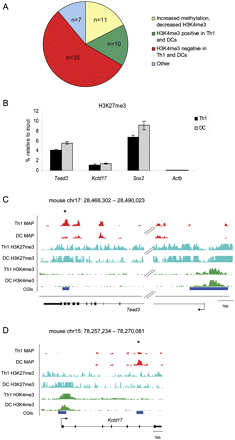
Most differentially methylated intragenic CGIs are depleted for H3K4me3 in immune cells. (A) Association of intragenic CGIs differentially methylated in Th1 and dendritic cells with H3K4me3 as assessed by ChIP-seq. (Yellow) CGIs where increased DNA methylation is associated with decreased H3K4me3 (n = 11). (Green) CGIs that are positive for H3K4me3 in both cell types despite a difference in methylation (n = 10). (Red) CGIs that lack H3K4me3 in both cell types (n = 35). (Blue) CGIs showing a nonsignificant change in H3K4me3 (n = 5) or where increased DNA methylation is associated with increased H3K4me3 (n = 2). (B) ChIP-PCR reveals enrichment for H3K27me3 at Tead3 and Kctd17 in Th1 and dendritic cells (DC) as well as at Sox2 (positive control). The active Actb gene acted as a negative control. These results were confirmed and extended using H3K27me3 ChIP-seq. MAP-seq (red), H3K27me3 ChIP-seq (cyan), and H3K4me3 (green) read density profiles for intragenic CGIs in the (C) Tead3 and (D) Kctd17 genes that lack H3K4me3 in Th1 and dendritic cells, despite showing differential methylation between the two cell types (asterisked CGI).











