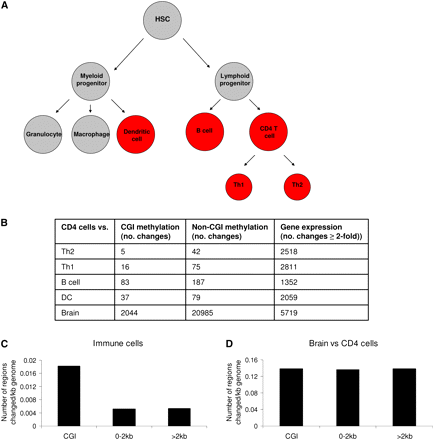
Cell type–specific methylation in the hematopoietic lineage detected by MAP-seq preferentially occurs at CGIs. (A) The immune cell lineage with the cell types investigated shown in red. HSC: hematopoietic stem cell. (B) The number of CGI methylation, non-CGI-associated methylation, and gene expression changes observed when CD4 T-cells were compared to Th2, Th1, B cells, dendritic cells (DC), and brain. (C,D) The location of methylation differences detected by MAP-seq. The fraction of the genome that can be interrogated by MAP-seq was categorized as overlapping a CGI (CGI), not overlapping but within 2 kb of a CGI (0–2 kb), or >2 kb from a CGI (>2 kb). The location of DNA methylation changes was then determined and expressed as the number of methylation changes occurring per kilobase (kb) of genome in each category. (C) Location of all DNA methylation changes occurring between different immune cells. (D) Location of DNA methylation changes occurring between brain and CD4 cells.











