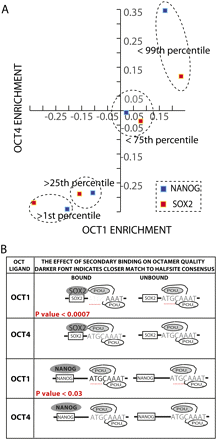
POU2F1(Oct1)/POU5F1(Oct4) binding is influenced by proximal SOX2 and NANOG binding. (A) Subsets of the oligonucleotide pool were created by ranking the pool by SOX2 (red squares) and NANOG (blue squares) enrichment. For each subset, a data point was plotted comparing the average POU2F1 (x-axis) and POU5F1 (y-axis) binding enrichment. (B) The top 1% of POU2F1 (Oct1) and POU5F1 (Oct4) enrichments are denoted as POU2F1 and POU5F1 “ligands” and annotated with an octamer binding model that allowed up to three mismatches. The number (n) and quality of octamer sites were compared for the POU ligands that did or did not bind SOX2. The darker font indicates closer agreement of a halfsite to the octamer consensus in the subset of POU ligands bound by SOX2. The lighter font indicates less agreement. The same analysis was repeated for NANOG. Statistically significant differences in comparisons of half-site strength are listed in red.











