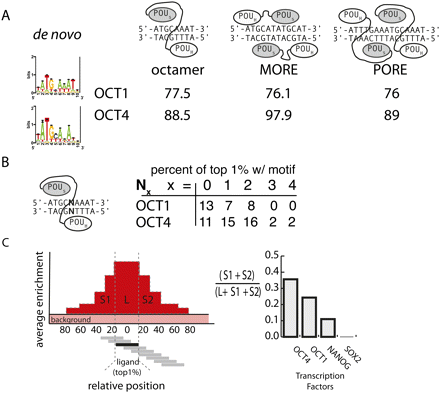
Identifying the specificity and clustering tendency of transcription factors. (A) De novo motif finding was used to identify sequence motifs enriched in the top 1% of oligonucleotides ranked by enrichment in the POU2F1 (Oct1) and POU5F1 (Oct4) immunoprecipitate. The average percentile rank of oligonucleotides matching the three existing POU factor binding models was recorded. Reported conformations of POU factor binding are shown. (B) The distribution of the top 1% of POU2F1 (Oct1) and POU5F1 (Oct4) ligands within gapped octamers is depicted. (C) The tendency of binding events to cluster was analyzed by measuring the average enrichment in the neighborhood of the top 1% of ligands. The percentage of area of the peak that fell outside of the ligand was used to measure a transcription factor's clustering tendency.











