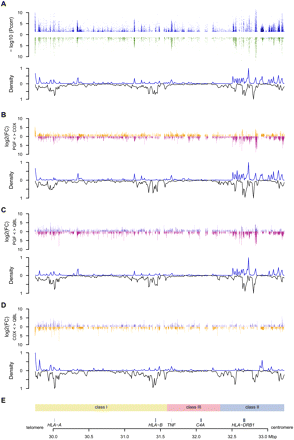
Distribution of differentially expressed (DE) probes versus polymorphic SNPs. Only probes shared by the three haplotypes were included. (A) Three-haplotype comparison. (Upper panel) Significance level of DE probes for either unstimulated (blue) or stimulated (green) cells. The −log10 of significant adjusted P-values are plotted against the genomic coordinates. (Lower panel) Density curve of DE probes normalized using the number of probes designed (upward) mirroring the density curve of polymorphic SNPs between the three cell lines (downward) for 350 10-kb windows spanning the MHC. Densities have been normalized. (B–D) Pairwise comparisons of COX versus PGF, QBL versus PGF, and QBL versus COX. For each pair, the log2 of the intensity fold change (FC) is represented in the upper panel. For example, when expression is higher in COX than in PGF, the FC is set positive and an orange bar is represented above the x-axis. Conversely, when expression is higher in PGF, the FC is negative and represented by a pink bar below the x-axis. The density curves of DE probes and of SNPs polymorphic between both cells are plotted in the lower panel. (E) Genomic context.











