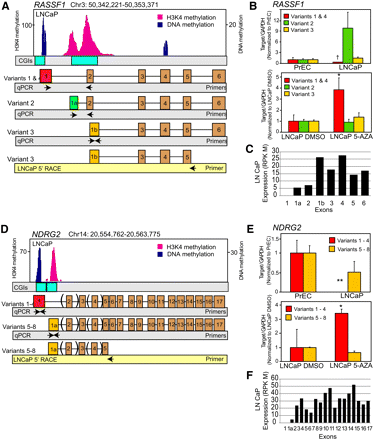
Regulation of alternate transcription start site utilization by DNA methylation. (A,D) Cancer-specific DNA methylation enables switching of alternative transcriptional start sites (TSS) leading to transcript isoform regulation. Next-generation sequencing for DNA methylation and histone 3 lysine 4 trimethylation (H3K4me3) in LNCaP cells reveals genome-wide patterning that couples CpG methylation with H3K4 marks to repress or activate specific transcript variants. Independent epigenetic modifications mark specific alternative TSS. In RASSF1 (A) and NDRG2 (D), CpG methylation occurs at the TSS of the longer variants, with H3K4me3 marks positioned on the TSS of the shorter variants. (B,E) Preferential silencing and 5-Aza-induced re-expression of CpG-methylated variants in LNCaP cells. Variants exhibiting CpG methylation on their TSSs show preferential silencing compared to variants with H3K4me3 marks in LNCaP cells. These variants show preferential reexpression upon treatment of LNCaP cells with 6 μM 5-Aza for 48 h. qRT-PCR data are normalized to variant expression levels in PrEC prostate primary epithelial cells or DMSO-treated LNCaP cells in the respective panels. (B,D) 5′-RACE results validated RASSF1 variant-3 and NDRG2 variants 5–8 expression in LNCaP cells. (C,F) Exon expression values from LNCaP RNA-seq data support the corresponding variant transcription of RASSF1 and NDRG2 genes.











