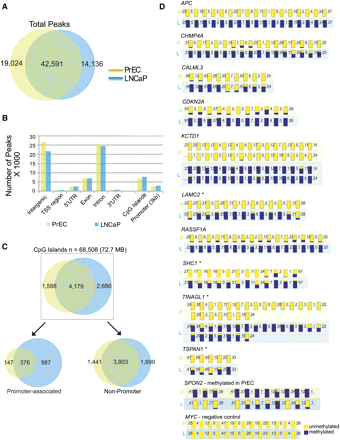
Characterization of genome-wide methylation patterns in prostate cells by M-NGS. (A) The Venn diagram represents a 70% overlap between the regions methylated in LNCaP (blue) and PrEC (green) cells. (B) In LNCaP (blue) and PrEC (green), the majority of DNA methylation occurred in intergenic and intronic regions, and the genomic distribution of methylation peaks was similar. (C) Promoter-associated CpG islands displayed a sevenfold difference in methylation between LNCaP (blue) and PrEC (green) cells. (D) DNA methylation in APC, CHMP4A, CALML3, CDKN2A, KCTD1, LAMC2, RASSF1, SHC1, TINAGL1, and TSPAN1 gene promoters in LNCaP (L) cells. SPON2 in PrEC (P) cells and a negative control region in MYC were validated by bisulfite sequencing. The methylation status of each CG residue from 10 clones sequenced on both strands was analyzed using the BIQ Analyzer (Bock et al. 2005) program, where the height of the blue bar indicates the percent methylation at a given position, yellow indicates no methylation, and the numbers indicate the distance between analyzed CG dinucleotides. *CpG islands were absent in these promoters. Additional details are provided in Supplemental Figures 5–7 and Supplemental Table 5. Validation of additional candidates, including NAP1L5, C9orf125, AOX1, AMT, NTN4, and PPP1R3C, are presented in Supplemental Figure 8.











