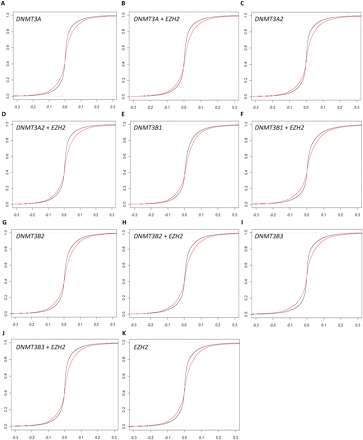
Overexpression of DNMTs and EZH2 results in increased methylation at a subset of prostate tumor hypermethylation sites. Ideal (black) and empirical (red) cumulative distribution functions of change in DNA methylation after DNMT or EZH2 transfection into cultured normal prostate cells. The empirical distribution functions are based on the 5912 CpGs that were hypermethylated in prostate tumors, while the ideal distribution functions are based on all 26,333 CpGs assayed on the array. Overexpression of (A) DNMT3A, (B) DNMT3A2, (C) DNMT3B1, (D) DNMT3B2, (E) DNMT3B3, (F) EZH2, (G) DNMT3A and EZH2, (H) DNMT3A2 and EZH2, (I) DNMT3B1 and EZH2, (J) DNMT3B2 and EZH2, and (K) DNMT3B3 and EZH2.











