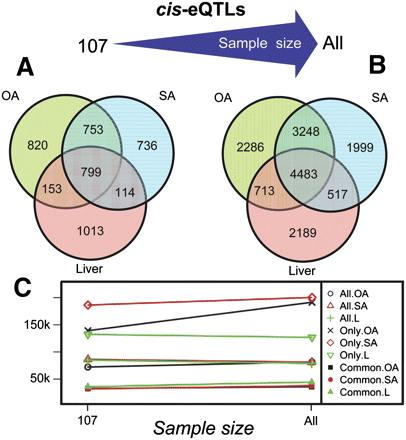
Cis-eQTLs overlaps. (A) Venn diagram for the cis eSNPs detected in three tissues using a subpopulation of 107 individuals. (B) Overlaps of the cis eSNPs identified in three tissues at maximum sample size in the RYGB cohort. Regardless of sample size, the OA and SA tissues share many more cis eQTLs in comparison to liver versus adipose. (C) Plot of mean distance of the eSNP to the gene. Symbols marked in the legend with “Only” represent the subset of the cis eSNPs specific to a particular tissue, “All” for all cis eSNPs detected in a tissue, and “Common” for the subset of cis eSNPs common to all three tissues. Cis eSNPs common to all three tissues are significantly closer to the associated genetic signal regardless of the sample size or tissue in which they were detected compared to cis eSNPs unique to a single tissue. The mean distance for all cis eQTLs detected in each tissue show little change between the small subpopulation and the full sample.











