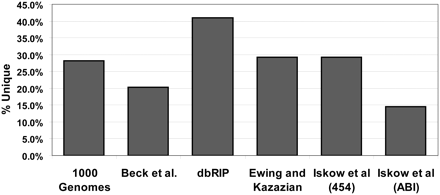
L1 insertions present in only one data set. The columns represent the fraction of insertions present in only the indicated data set out of all validated insertions in the data set (unique). Insertions can be validated by site-specific PCR, sequencing spanning the element, or presence in another independent data set, depending on the study. Iskow et al. (2010) presented data generated using two different sequencing methods as noted in the column labels. We analyzed the paired-end Illumina data from the 1000 Genomes Project (The 1000 Genomes Project Consortium 2010), Beck et al. (2010) employed a fosmid-end resequencing strategy, dbRIP cross-references data sets generated using a wide variety of techniques (Wang et al. 2006), and Ewing and Kazazian (2010) used Illumina sequencing.











