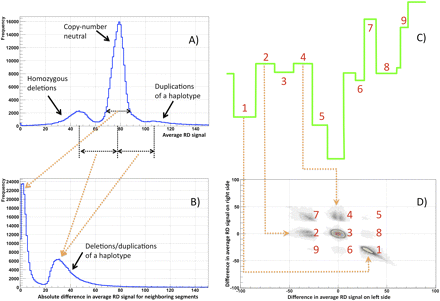
Statistics on partitioning the RD signal for a child in a CEPH trio (NA12878) using 100-bp bins and standard parameters (see Methods). (A) Average RD signal distribution in produced segments. The distribution has three clear peaks: around the genomic RD average (no CNVs), half of that (heterozygous deletion), and one and one-half of that (duplication of one haplotype). The average genomic RD signal is ∼ 77 reads. Not all partitioned regions with abnormal RD are called CNVs by the statistical significance test. Therefore, the area under each peak is not representative of the corresponding fraction of CNVs. (B) Distribution of the average RD signal difference for neighboring segments. The distribution is for the absolute value of the difference and shows that either produced segments have similar average signals (peak around zero) or their average signals are approximately half of the genomic average RD signal (second peak), indicating deletion/duplication of one haplotype. (C) Example of partitioning clarifying clusters in D. (D) Distribution of the average RD signal difference at the left and right boundary for each segment. The distribution has several clear clusters. Clusters originate due to various combinations of segments with different RD signals. Clusters 8 and 9 represent cases of enclosed events, such as duplication of a region within duplication.











