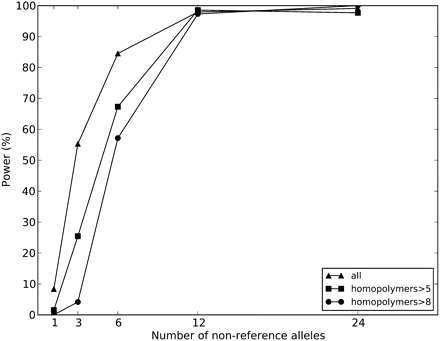
Power as a function of non-reference allele frequency in a simulated pooled analysis. The Dindel Bayesian EM algorithm was used to detect indels in a pool of 60 individuals with simulated read-depth of 4×. Due to increased sequencing error indel rate in long homopolymers, power decreases as a function of homopolymer length. Calls were made using a 99% confidence threshold on the posterior probability of a non-reference indel variant being present.











