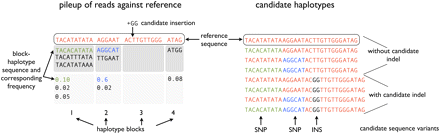
Procedure for generation of candidate haplotypes. We first consider the empirical distribution of bases determined from the initial alignments of reads to the reference and infer a heuristic haplotype block model to preserve sequences that always occur together in one read. We then choose n block-haplotypes with the highest empirical frequency, and generate candidate haplotypes by considering all combinations of these n block-haplotypes. The number of candidate haplotypes obtained this way is thus 2n. It is possible that multiple subhaplotypes from the same block are chosen. In the second step, all candidate variants (most importantly, the candidate indels) are added to these n candidate haplotypes, resulting in a set of, at most,k · 2n candidate haplotypes, where k is the number of candidate variants tested.











