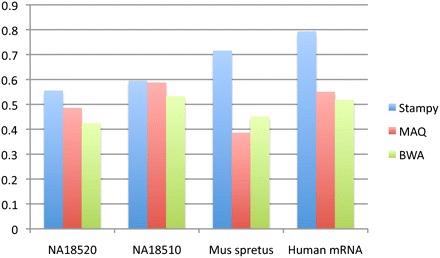
Pairwise concordance of independently mapped reads. The data (two human samples from the 1000 Genomes Project (The 1000 Genomes Project Consortium 2010); a divergent mouse subspecies; and human mRNA from an MCF-7 cell line; see text) were mapped to the human or mouse reference genomes (both NCBI build 37) by considering each read of a pair independently. Concordance was calculated as the proportion of reads that mapped to within 500 bp (for genomic DNA) or 10,000 bp (for the mRNA data set) of its mate.











