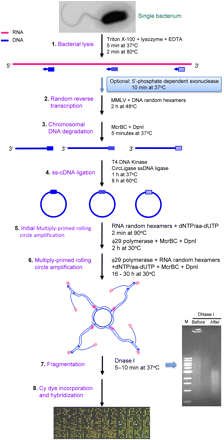
Single-bacterium total transcript amplification strategy. This strategy is developed based on multiply primed rolling circle amplification of circularized cDNA using ϕ29 DNA polymerase. Blue boxes indicate DNA random hexamers, and pink boxes indicate RNA thiophosphate-linked random hexamers. The purpose and components of each step are indicated (for further details, see text). Abbreviations are as follows: aa-dUTP indicates 5-(3-aminoallyl)-2′-deoxyuridine-5′-triphosphate; dNTP, deoxyribonucleotide triphosphate; DpnI, restriction endonuclease from Diplococcus pneumoniae; EDTA, ethylenediaminetetraacetic acid; ϕ29 polymerase, DNA polymerase from a Bacillus subtilis phage Phi29; M, 1 kb DNA marker from New England BioLabs; McrBC, E. coli homing endonuclease; MMLV, Moloney murine leukemia virus reverse transcriptase; and ss-cDNA, single-stranded cDNA.











