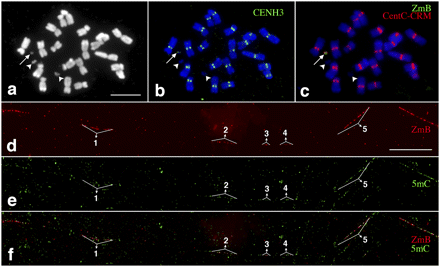
Mapping of DNA methylation in a significantly rearranged minichromosome derived from the maize B chromosome. (a) A metaphase cell derived from a maize line containing a mini-B chromosome (arrow). Two arrowheads point to the two satellites associated with maize chromosome 6. Bar, 5 μm. (b) Immunofluorescence assay using anti-CENH3 antibody (green). The arrow points to the CENH3 signal associated with the minichromosome. (c) FISH mapping of CentC-CRM (red) and ZmB (green) repeats. The arrow points to the signals associated with the minichromosome. (d) Fiber-FISH signals from the ZmB repeat located in the mini-B chromosome. Bar, 20 μm. (e) Detection of 5mC on the same DNA fiber as d. (f) A merged image of the ZmB and 5mC signals. The ZmB signals were all associated with 5mC signals on the DNA fiber. The five ZmB repeat arrays associated with the CENH3 domain are labeled in d–f.











