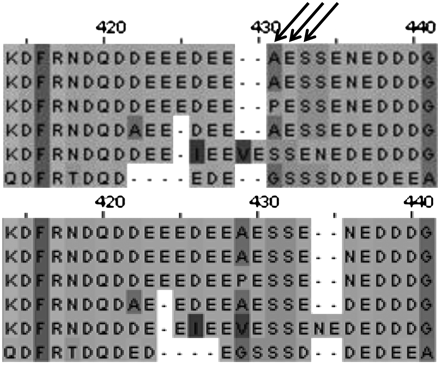
Figure 3.
Example alignment from the genes that we performed visual inspection on. The illustrated alignment segment is from the melanogaster group alignments of gene FBgn0036686. The top alignment was made with T-Coffee and has three sites inferred under selection at 99% cutoff. The bottom alignment was made with ProbCons; the highest posterior probability in this region for the ProbCons alignment is 60%.











