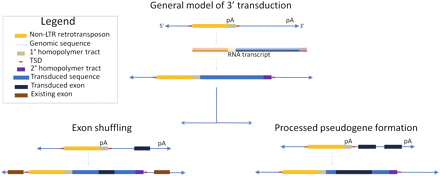
Schematic illustrating the mechanism of 3′ transduction by non-LTR retrotransposons and possible gene-related impacts. TE-mediated 3′ transduction occurs when the transcription machinery skips a weak or nonexistent polyadenylation signal (pA). Transcription continues until a downstream polyadenylation signal is recognized. The resulting transcript will contain a portion of the 3′ genomic flank and a secondary homopolymer tract, which will be reverse transcribed into cDNA upon reinsertion into the genome (Boeke and Pickeral 1999; Moran et al. 1999; Goodier et al. 2000). If the transduced sequence contains an exon, it may be inserted near existing exons, resulting in an exon shuffling event. Assuming RNA pol II transcription and normal post-transcriptional processing, two or more exons in the transduced sequence may be merged and reinserted, resulting in a processed pseudogene.











