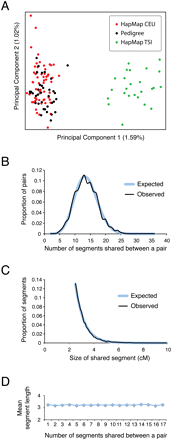
Characteristics of HapMap CEU parents as a background reference population. (A) Principal Components Analysis comparing 36 individuals from our three pedigrees (no pair closer than seventh-degree relatives) to 85 unrelated individuals from three European populations (60 HapMap CEU parents from 30 parent–offspring trios and 25 HapMap TSI individuals) based on pairwise allele-sharing distances computed from ∼247,000 SNPs typed on the Affymetrix SNP array (for additional details on data and methods, see Xing et al. 2010). The percentage of genetic variation explained by each component is given on the corresponding axis. (B) Distribution of the number of segments with length >2.5 cM that are inferred to be shared IBD by Germline in pairs of CEU individuals (Observed), with fitted Poisson distribution (Expected). (C) Distribution of the lengths of IBD segments longer than 2.5 cM in CEU pairs (Observed), with fitted exponential distribution (Expected). (D) Scatterplot of the number of IBD segments per pair versus the mean length of segments in the pair.











