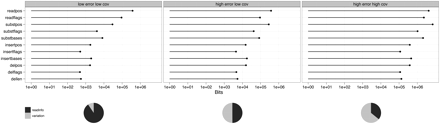
Storage components for three parameterizations of simulated data: 0.1% error and 1× coverage (left panel), 1% error and 1× coverage (middle), and 1% error and 25× coverage (right). readpos and readflags is the storage of the read positions and read flags (strand, exact match), respectively. Variation storage for substitutions (subst), insertions (insert), and deletions (del) is split into positional information (pos), flags (flags), and bases (bases, for substitutions and insertions) or length (len, for deletions). The pie charts show overall storage requirements, where readinfo sums over read positions and read flags, and variation is the sum over all variation storage components.











