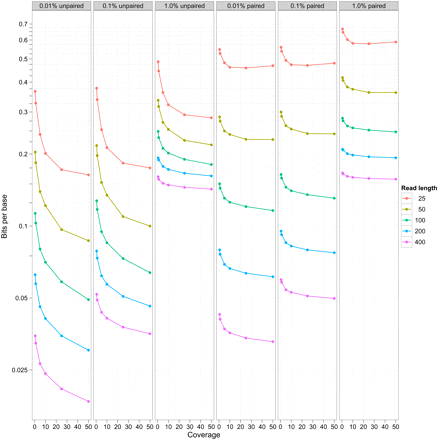
Compression efficiency for simulated data sets. The plot shows storage of DNA sequence expressed as a bits/base stored on the y-axis (log scale) vs. coverage of data sets (x-axis) for different read lengths (the different colors) after reference-based compression. The different columns indicate different simulated error rates (0.01%, 0.1%, 1.0%). The left three panels show this for unpaired data, the right three for paired data.











