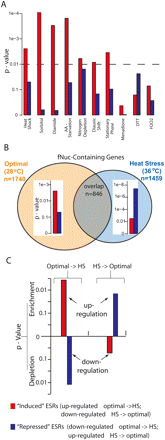
Fragile nucleosomes in the promoters of the ESR gene correlate with transcription up-regulation and are inclined to be evicted upon environment changes. (A) Bar graphs depict the P-values of the enrichment of subsets of the “induced” ESR genes that are up-regulated (red) or the “repressed” ESR genes that are down-regulated (blue) in response to specific environmental stresses in fragile nucleosome-containing genes. (Dashed line) The P-value cutoff at 0.01. (B) The spectra of fragile nucleosome-containing genes identified under optimal growth (28°C; light brown) or heat stress (36°C; light blue) are significantly different. Inserted bar diagrams illustrate the P-values of enrichment of the “induced” or the “repressed” subgroups of ESR genes (red and blue) in corresponding fragile nucleosome-gene groups. Identifications of ESR subgroups and their transcription changes with temperature shifts are obtained from Gasch et al. (2000). (C) Enrichment of fragile nucleosomes that are converted to nucleosome-free regions in combinations of ESR subgroup plus environment change that lead to transcription up-regulation. The y-axis represents the log-scale P-values calculated by comparing the number of fNuc to NFR conversions in the “induced” or “repressed” ESR genes with that in all genes. Positive and negative P-values indicate enrichment and depletion, respectively.











