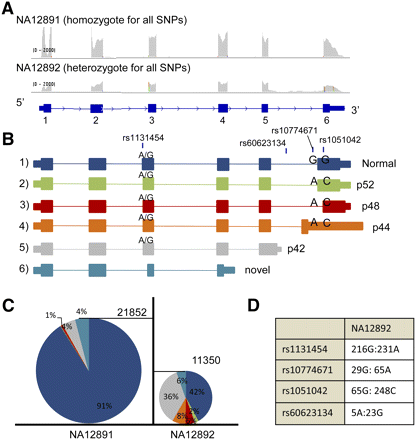
OAS1 isoforms. (A) Screenshots from IGV displaying the read coverage within the OAS1 gene. The maximum height of both tracks is 2000. (B) All possible isoforms seen in OAS1 and their association with SNPs rs1131454, rs60623134, rs10774671, and rs1051042. Larger font indicates stronger allelic imbalance (not to scale), and if both alleles are shown, then there is no evidence for genetic control on isoform production by this SNP. Isoforms 1, 2, 3, and 4 differ only by exon 6: in isoform 1 it is the normal exon 6; in isoform 2 it is shifted by 1 bp (exon 6′); in isoform 3 it is shifted by 98 bp (exon 6″); and in isoform 4 it begins within the intron (exon 6″′). (C) Relative percentages of isoforms seen within individuals NA12891 and NA12892. The number of isoforms is inferred by the number of reads mapping to unique splice junctions for most isoforms (junction 5-6 for isoform 1; junction 5-6′ for isoform2; junction 5-6″ for isoform 3; junction 5 to 6″′ for isoform 4; junction 2-3′ for isoform 6). For isoform 5, expression was measured by the number of reads mapping 25 bp past the end of exon 5 (still within the extended exon). The height of each pie chart is representative of the gene expression for OAS1. There is a 1.9-fold increase in expression in favor of individual NA12892. (D) The number of reads supporting the reference and alternate alleles seen in individual NA12892.











