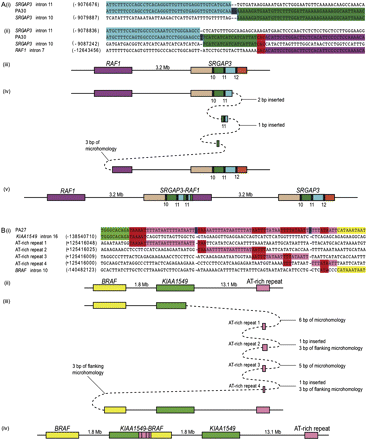
Breakpoint analyses of two complex rearrangements. The figure is not to scale. In both panels, microhomology (red); insertions (blue). Genes are depicted by their orientation on the minus strand. (A) Complex rearrangement in PA30. (Purple) RAF1; (beige) SRGAP3 intron 9; (green) intron 10; (light blue) intron 11; (orange) intron 12. Exons are numbered and represented by black rectangles. (i) Sequence alignment showing the SRGAP3 intron 11–intron 10 breakpoint. (ii) Sequence alignment for the SRGAP3–RAF1 breakpoint. (iii) Schematic representation of the region on chromosome 3p25 prior to duplication. (iv) The three template switching events that may have given rise to the complex breakpoint in PA30. (v) Observed complex rearrangement. (B) Complex rearrangement in PA27. (Light green) KIAA1549; (yellow) BRAF; (pink) AT-rich region. (i) Sequence alignment showing the KIAA1549–BRAF breakpoint in PA27. (ii) Schematic representation of the region on chromosome 7q34 prior to duplication. (iii) The five template switching events that could have generated the complex breakpoint in PA27. (iv) Observed complex rearrangement.











